Abstract
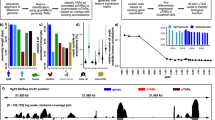
Transcriptional profiling is a powerful approach for studying mouse development, physiology and disease models. Here we describe a protocol for mouse thiouracil tagging (TU tagging), a transcriptome analysis technology that includes in vivo covalent labeling, purification and analysis of cell type–specific RNA. TU tagging enables the isolation of RNA from a given cell population of a complex tissue, avoiding transcriptional changes induced by cell isolation trauma, as well as the identification of actively transcribed RNAs and not preexisting transcripts. Therefore, in contrast to other cell-specific transcriptional profiling methods based on the purification of tagged ribosomes or nuclei, TU tagging provides a direct examination of transcriptional regulation. We describe how to (i) deliver 4-thiouracil to transgenic mice to thio-label cell lineage–specific transcripts, (ii) purify TU-tagged RNA and prepare libraries for Illumina sequencing and (iii) follow a straightforward bioinformatics workflow to identify cell type–enriched or differentially expressed genes. Tissue containing TU-tagged RNA can be obtained in 1 d, RNA-seq libraries can be generated within 2 d and, after sequencing, an initial bioinformatics analysis can be completed in 1 additional day.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Gay, L. et al. Mouse TU tagging: a chemical/genetic intersectional method for purifying cell type-specific nascent RNA. Genes Dev. 27, 98–115 (2013).
Miller, M., Robinson, K., Cleary, M. & Doe, C. TU-tagging: cell type–specific RNA isolation from intact complex tissues. Nat. Methods 6, 439–441 (2009).
Cleary, M.D., Meiering, C.D., Jan, E., Guymon, R. & Boothroyd, J.C. Biosynthetic labeling of RNA with uracil phosphoribosyltransferase allows cell-specific microarray analysis of mRNA synthesis and decay. Nat. Biotechnol. 23, 232–237 (2005).
Friedel, C.C. & Dölken, L. Metabolic tagging and purification of nascent RNA: implications for transcriptomics. Mol. Biosystems 5, 1271–1278 (2009).
Lai, S.L., Miller, M.R., Robinson, K.J. & Doe, C.Q. The Snail family member Worniu is continuously required in neuroblasts to prevent Elav-induced premature differentiation. Dev Cell 23, 849–857 (2012).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Doyle, J.P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135, 749–762 (2008).
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135, 738–748 (2008).
Deal, R.B. & Henikoff, S. The INTACT method for cell type-specific gene expression and chromatin profiling in Arabidopsis thaliana. Nat. Protoc. 6, 56–68 (2011).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl. Acad. Sci. USA 106, 13939–13944 (2009).
Kenzelmann, M. et al. Microarray analysis of newly synthesized RNA in cells and animals. Proc. Natl. Acad. Sci. USA 104, 6164–6169 (2007).
Feil, R. et al. Ligand-activated site-specific recombination in mice. Proc. Natl. Acad. Sci. USA 93, 10887–10890 (1996).
Diez-Roux, G. et al. A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol. 9, e1000582 (2011).
Adiconis, X. et al. Comparative analysis of RNA sequencing methods for degraded or low-input samples. Nat. Methods 10, 623–629 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics (Oxford, England) 25, 2078–2079 (2009).
Anders, S. HTSeq: Analyzing high-throughput sequencing data with Python http://www-huber.embl.de/users/anders/HTSeq/doc/overview.html(2011).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Acknowledgements
We thank the University of Oregon (UO) Genomics Core Facility for Illumina sequencing; V. Devasthali, U. Hostick and the UO Transgenic Mouse Facility for support generating the CA>GFPstop>HA-UPRT mice; P. Batzel for critical input on the bioinformatics workflow; and B. Simek for beta-testing the bioinformatics protocol. The use of the UO's applied computational instrument for scientific synthesis (ACISS) server was supported by a Major Research Instrumentation grant from the National Science Foundation (no. OCI-0960354). Funding was provided by The US National Institutes of Health (NIH) grant nos. NIH 5R00HL087598 (K.S.) and NIH 3R01DE013085 (Kaartinen, V.S.), and by the Howard Hughes Medical Institute (C.Q.D.).
Author information
Authors and Affiliations
Contributions
L.G., K.V.K., M.R.M., C.Q.D. and K.S. developed the protocol. L.G., K.V.K., C.Q.D. and K.S. prepared and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Example counts table used in DESeq. The data is for a Type I experiment determining the Tie2:Cre-lineage transcriptome of P6 brains. Data is provided for two pairs of total and TU-tagged RNA samples. (TXT 415 kb)
Supplementary Table 2
Example of DESeq output showing the top 20 most Tie2:Cre-lineage enriched transcripts in P6 whole brain (expression level filtered). The nine enriched positive control endothelial transcripts (of thirteen used) are also shown. (PDF 59 kb)
Supplementary Table 3
Complete DESeq output from the example Type I experiment defining the P6 brain Tie2:Cre lineage transcriptome. (TXT 275 kb)
Rights and permissions
About this article
Cite this article
Gay, L., Karfilis, K., Miller, M. et al. Applying thiouracil tagging to mouse transcriptome analysis. Nat Protoc 9, 410–420 (2014). https://doi.org/10.1038/nprot.2014.023
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.023
This article is cited by
-
Cell cycle specific, differentially tagged ribosomal proteins to measure phase specific transcriptomes from asynchronously cycling cells
Scientific Reports (2024)
-
Sequencing cell-type-specific transcriptomes with SLAM-ITseq
Nature Protocols (2019)
-
Genome-wide real-time in vivo transcriptional dynamics during Plasmodium falciparum blood-stage development
Nature Communications (2018)
-
“Hit-and-Run” transcription: de novo transcription initiated by a transient bZIP1 “hit” persists after the “run”
BMC Genomics (2016)
-
Identification of sensory hair-cell transcripts by thiouracil-tagging in zebrafish
BMC Genomics (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.