Abstract
Aim:
To develop a C. elegans model of amyotrophic lateral sclerosis (ALS) and to evaluate the role of autophagy in the disease.
Methods:
Stable transgenic worms expressing the G93A mutant form of Cu,Zn-superoxide dismutase (SOD1) in GABAergic motor neurons were generated. Axon guidance and protein aggregation in the motor neurons were observed with fluorescence microscopy. A paralysis assay was performed to evaluate the motor function of the transgenic worms. The expression of autophagic genes in daf-2(e1370) mutants was analyzed using real-time PCR. The reporter GFP::LGG-1 was used to verify whether autophagy was induced in motor neurons.
Results:
Expression of G93A SOD1 in motor neurons caused age-dependent motor defects accompanied by significant SOD1 aggregation and axon guidance failure. After 12 d, over 80% of the G93A worms became paralyzed, whereas less than 10% of the controls showed a paralytic phenotype. In the daf-2(e1370) mutants of C. elegans, the levels of autophagic genes bec-1, atg-7, lgg-1, and atg-18 were upregulated by approximately 1.5-fold, the level of unc-51 increased by approximately fourfold, and autophagosomes in motor neurons was markedly increased. Crossing the daf-2(e1370) mutation into the G93A SOD1 mutant worms significantly ameliorated the motor defects, SOD1 aggregation, and axon guidance failure.
Conclusion:
G93A SOD1 expression in motor neurons of C. elegans results in characteristic alterations of ALS. Increased autophagy protects C. elegans motor neurons against the toxicity of mutant SOD1.
Similar content being viewed by others
Introduction
Autophagy is a ubiquitous intracellular catabolic system for bulk degradation. It is a response to cell stressors, such as damaged organelles or aggregate-prone toxic protein, that prevents their detrimental effect1. Autophagy begins with the formation of a double-membrane structure, termed the autophagosome, which engulfs the cytoplasm and/or organelles and then subsequently fuses with a late endosome or lysosome to form an autophagolysosome. Inside the autophagolysosome, lysosomal hydrolases degrade the sequestered materials2. Autophagy can be upregulated by starvation and can also be triggered by protein aggregates, oxidative stress, and damaged cytoplasmic organelles3. It plays an important role in many physiological processes, including the response to nutrient stress, innate and adaptive immunity, and autophagic cell death4,5. Autophagy also functions as an important quality control system by eliminating disease-related mutant proteins that are associated with various neurodegenerative disorders such as Alzheimer's disease, Parkinson's disease, Huntington's disease, and amyotrophic lateral sclerosis (ALS)6,7,8,9,10.
ALS is a late-onset, fatal paralytic disorder characterized by the death of motor neurons in the brain and spinal cord. Approximately 10% of ALS cases are familial ALS (FALS)11,12. The first gene to be identified as associated with FALS was SOD113,14. Mutations in the SOD1 gene cause the most prevalent form of FALS, accounting for approximately 20% of all FALS cases. More than 160 different mutations have been identified in SOD1-associated FALS cases15. Mutations in SOD1 usually cause a dominant gain of function rather than a loss of dismutase activity16. Mice that overexpress mutant SOD1 develop a severe motor neuron disease, the symptoms of which is similar to that exhibited by ALS patients10,17,18. Revealing the mechanism of motor neuron dysfunction caused by mutant SOD1 may help in understanding the progression of ALS in patients and in finding a potential treatment for ALS. It has been widely accepted that motor neuron degeneration is associated with the misfolding and aggregation of SOD112. However, the exact mechanism whereby mutant SOD1 leads to the dysfunction and death of motor neurons remains elusive. In addition, it is important to investigate the role of protein degradation pathways, especially the autophagy system, in the degradation of mutant SOD1 in motor neurons.
It has been reported that autophagy contributes to the degradation of both wild-type and mutant SOD1 in neural cell lines19. In addition, increased autophagy has been observed in G93A SOD1 transgenic mice20,21. Increasing autophagy by knocking out X-box-binding protein-1 enhances the clearance of SOD1 aggregates and delays the progression of the disease in G86R SOD1 transgenic mice22. The pharmacological agent lithium, which can enhance autophagy, delays the progression of ALS in both transgenic ALS mice and ALS patients10. Therefore, autophagy is expected to have a beneficial effect in ALS. However, another autophagy enhancer, rapamycin, was found to accelerate motor neuron degeneration and shortened the lifespan of transgenic ALS mice23.
Here, we used Caenorhabditis elegans (C. elegans) as a model organism to address whether autophagy has a beneficial or detrimental effect on ALS in vivo. Conservation of genetic and metabolic pathways between C. elegans and mammals, as well as the visualization of neurons in vivo and the relative ease with which genetic manipulations can be performed, make C. elegans an excellent model for neurodegenerative diseases. The nervous system of C. elegans is composed of 302 neurons, which utilize most of the known neurotransmitters in the mammalian nervous system, including GABA, dopamine, glutamate, serotonin, and acetylcholine. C. elegans has been utilized in modeling various neurodegenerative diseases, including polyglutamine expansion diseases, α-synuclein-linked Parkinson disease, and Aβ-associated Alzheimer's disease24,25,26. Previous studies have used C. elegans that express G85R SOD1 pan-neuronally as an ALS model27,28.
Because mutant SOD1 mainly destroys motor neurons in ALS patients, we made a transgenic C. elegans that was engineered to express human G93A SOD1 in motor neurons and investigated the specific effect of mutant SOD1 on motor neurons. We found that G93A SOD1 transgenic worms developed a significant motor dysfunction that was associated with the aggregation of SOD1 in motor neurons and the loss of axons that project to the dorsal nerve cord. The daf-2 gene encodes a receptor tyrosine kinase that is the C. elegans insulin/IGF receptor ortholog, and the loss-of-function mutant daf-2(e1370) increases autophagy in hypodermal seam cells29,30. Here, we present evidence that the daf-2(e1370) mutant shows increased autophagic gene expression. Furthermore, we found that motor neurons of the daf-2(e1370) mutant show increased autophagy, which could protect against G93A SOD1-induced motor defects in C. elegans.
Materials and methods
Molecular biology
The unc-25 promoter was amplified by KOD-PLUS Neo DNA polymerase from N2 genomic DNA. Then, the unc-25 promoter was inserted into the HindIII and BamHI sites of pPD95.77 to generate pPD95.77-unc-25::GFP. The 465 bp human G93A SOD1 was inserted downstream of the Not I site to generate pPD95.77-unc-25::G93A SOD1::GFP. The 465 bp human G93A SOD1 with a TAA stop code was inserted downstream of the NotI site to generate pPD95.77-unc-25::G93A SOD1. The autophagy reporter unc-47::GFP::LGG-1 was constructed by inserting the 1194 bp unc-47 promoter and the coding sequence of lgg-1 into the plasmid pPD117.01.
Transgenic C. elegans
Extrachromosomal arrays were attained via injection into the germline using standard techniques31. For GFP, G93A SOD1-GFP, or G93A SDO1 expression, 100 ng/μL DNA was injected into the adult gonad. The unc-47::GFP::LGG-1 plasmid was injected at 20 ng/μL. The G93A SOD1 transgene was integrated into the genome by exposing worms to 4,5′,8-trimethylpsoralen combined with UV light, thus generating stable transgenic lines. Three lines were selected and then outcrossed five times with N2 worms.
C. elegans strains
Worms of the Bristol strain N2 were used as wild-types. N2 worms and daf-2(e1370) mutants were obtained from the Caenorhabditis Genetics Center, which is funded by the NIH National Center for Research Resources. Experiments were performed at 20 °C using standard C. elegans techniques32.
Paralysis analysis
Worms were scored as paralyzed if they moved their noses but failed to move their bodies when their noses were tapped with a platinum worm picker. Experiments were performed with over 20 worms per plate in triplicate.
Fluorescence microscopy
Worms were immobilized in 5 mmol/L sodium azide in M9 buffer on 2% agar pad slides. Images were collected with a Leica TCS SP5 confocal laser scanning microscope. To analyze motor neuron axon guidance defects in C. elegans, the number of DD/VD axons that reached the dorsal nerve cord in either L4 larvae or young adults were counted. At least 20 worms of each strain were examined.
Quantitative real-time PCR and primers
Total RNA was isolated from worms using TRIzol reagent. Reverse transcriptase was used for oligo (dT)-primed first-strand cDNA synthesis. We performed real-time PCR to quantify the mRNA levels of autophagic genes. PCR amplification was performed using an Applied Biosystems 7500 Real-Time PCR System in a mixture of SYBR Green Real-time PCR Master Mix and 0.4 mmol/L aliquots of each primer in a final volume of 20 μL. Expression levels relative to the wild-type N2 strain were normalized to two endogenous reference genes (act-1 and ama-1) and were calculated using Applied Biosystems 7500 software V2.0.5.
Real-time PCR primers designed for the C. elegans genes: ama-1 (forward primer: 5′-CGGTCAGAAAGGCTATCGAG-3′; reverse primer: 5′-CCAACCTCCTGACGATTGAT-3′), act-1 (forward primer: 5′-GCTGGACGTGATCTTACTGATTACC-3′; reverse primer: 5′-GTAGCAGAGCTTCTCCTTGATGTC-3′), unc-51 (forward primer: 5′-CGCCGGTGGTTCAGCGGATT-3′; reverse primer: 5′-TATCCTGGGTGTCGGCGGGG-3′), bec-1 (forward primer: 5′-ACGAGCTTCATTCGCTGGAA-3′; reverse primer: 5′-TTCGTGATGTTGTACGCCGA-3′), atg-18 (forward primer: 5′-CAGGAGCCGCAAGGAGTAAT-3′; reverse primer: 5′-CGATTGGTTGCTTGCTTCGG-3′), atg-7 (forward primer: 5′-CCAAAAGCTGTGGGATGGGA-3′; reverse primer: 5′-GCGTTCCAGCACCAAGAATG-3′), lgg-1 (forward primer: 5′-GCCGAAGGAGACAAGATCCG-3′; reverse primer: 5′-GGTCCTGGTAGAGTTGTCCC-3′).
Statistical analysis
All values are presented as the mean±SEM. A data analysis was performed using a two-way ANOVA or a t-test using GraphPad Prism5 version 5.02 software.
Results
Transgenic worms expressing human G93A SOD1 in motor neurons display age-dependent paralysis
To generate the G93A SOD1 transgenic worms, we used the unc-25 promoter to express GFP-tagged G93A SOD1 specifically in the 26 GABAergic motor neurons. GFP control worms were also generated using the same promotes (Figure 1A, 1B). The GFP tag does not affect the SOD1-induced phenotype. It has also been reported that SOD1-YFP and SOD1-GFP fusion proteins exhibit similar behavior to their non-fused counterparts in the context of producing ALS-like disease27,28,33,34. Therefore, for the convenience of directly observing SOD1 in motor neurons, we made stable transgenic lines of the GFP-fused G93A SOD1 worms (hereafter referred to as G93A worms). During adulthood, the transgenic worms began to exhibit an uncoordinated motility phenotype that progressed to paralysis. We performed a widely used paralysis assay to study the age-dependent degenerative phenotype of G93A worms25 and found that paralysis was age-dependent and occurred at a much higher rate in G93A worms compared with the GFP control worms (GFP expressed only in motor neurons) and N2 non-transgenic control worms (Figure 1E). After 12 d on plates, over 80% of the G93A worms became paralyzed, whereas less than 10% of the controls showed a paralytic phenotype, indicating that G93A SOD1 exhibits toxic properties in motor neurons and is sufficient to cause motor defects in C. elegans.
Motor defect in G93A worms. (A) Control worm that with GFP expression in the motor neurons shows normal axon guidance to the dorsal side. Scale bar: 100 μm. (B) Aggregation is not observed in the motor neuron of GFP control worms. Scale bar: 10 μm. (C) Axons of motor neurons fail to reach the dorsal side in G93A worms. Arrow: axon that fails to reach the dorsal side. Scale bar: 10 μm. (D) G93A SOD1-GFP aggregations in the motor neurons of G93A worms. Arrow: aggregation in cell body. Arrow head: aggregation in axon. Scale bar: 10 μm. (E) G93A worms show age-dependant paralysis. (F) Quantification of the number of axons that reach dorsal nerve cord in the control worms and G93A worms. G93A worms show significantly decreased number of axons that extend to dorsal side. (G) Quantification of the percentage of the motor neurons with aggregations in G93A worms over time. Mean±SEM. n=25. cP<0.01 vs GFP.
G93A SOD1 aggregates in neural cell bodies and causes axon guidance defects
It is known that toxic protein-induced neuronal dysfunction is often associated with protein aggregation. Abnormal accumulation of SOD1 inclusions in spinal motor neurons is a pathological hallmark of mutant SOD1-related FALS35. Fluorescence microscopy of G93A worms revealed that G93A SOD1 accumulated in the cell bodies of motor neurons, whereas the control worms showed no aggregations (Figure 1B, 1D). The aggregated SOD1 appeared as early as the L1 stage, and the aggregates increased in number with age (Figure 1G). The aggregations were also observed within the axons of the motor neurons (Figure 1D). In C. elegans, motor neurons are located along the ventral nerve cord and project axons to the dorsal nerve cord to innervate the dorsal muscle36 (Figure 1A). Compared with the control worms, the G93A worms exhibited axon guidance defects, as most of the axons failed to reach their dorsal destinations (Figure 1F), indicating that the damaged axon guidance caused by G93A SOD1 and the formation of SOD1 aggregates might contribute to the motor defects in G93A worms. In conclusion, we established a C. elegans model of FALS that showed motor dysfunction associated with axon guidance defects and SOD1 aggregation in motor neurons.
The daf-2(e1370) mutant shows increased autophagy
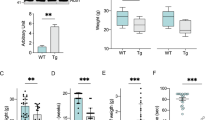
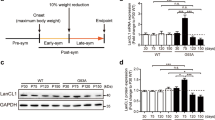
Next, we tried to find a C. elegans autophagic mutant to determine the role of autophagy in our C. elegans model of FALS. Previous studies of autophagy mainly rely on pharmacological agents such as lithium and rapamycin to enhance autophagy and detect its role in some ALS models. Because of the side effects of the pharmacological agents, it is important to find genetic tools to alter autophagy in vivo. In C. elegans, the loss-of-function mutation of daf-2, daf-2(e1370), increases autophagy29,30. Increased autophagy has been detected (both by visualization of GFP::LGG-1 puncta and by electron microscopy) in the hypodermal seam cells30. We performed real-time PCR to determine mRNA levels of autophagic genes in the daf-2(e1370) mutants. Autophagic function has been documented for genes that act in induction (unc-51/ATG1), vesicle nucleation (bec-1/ATG6, vps-34/VPS34), protein conjugation systems (atg-7/ATG7, lgg-1/ATG8, lgg-3/ATG12), retrieval and vesicle recycling (atg-18/ATG18) in the worms37. We chose unc-51, bec-1, atg-7, lgg-1, and atg-18 and analyzed the mRNA level of these genes in the daf-2(e1370) mutants. We found increased mRNA levels of these genes compared with wild-type N2 worms (Figure 2A). The levels of bec-1, atg-7, lgg-1, and atg-18 were upregulated by approximately 1.5-fold, whereas the level of unc-51 increased by approximately fourfold. These data indicate that the autophagic pathway is induced or activated in the daf-2(e1370) mutants. We also expressed the reporter GFP::LGG-1, which indicates formation of the autophagosome, in motor neurons, using the unc-47 promoter. We observed an increase in the number of GFP::LGG-1 puncta in motor neurons (Figure 2B, 2C), suggesting that autophagy is increased in motor neurons of the daf-2(e1370) mutant.
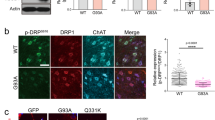
Autophagy is increased in the daf-2(e1370) mutant. (A) mRNA levels of autophagic genes detected by real-time PCR in wild-type N2 worms and daf-2(e1370) worms. All real-time PCR experiments were performed three times in duplicate. (B) GFP::LGG-1 expression in the motor neurons of wild-type N2, daf-2(e1370), and daf-2(e1370);G93A SOD1 (without GFP tag) L4 worms. The arrow shows representative small GFP::LGG-1 aggregate that labels autophagosomal structures. Scale bar: 5 μm. (C) Quantification of GFP::LGG-1–positive punctuate in the motor neuron of wild-type N2, daf-2(e1370), and daf-2(e1370);G93A SOD1 L4 worms (at least 50 motor neurons from 10 different worms were analyzed). Mean±SEM. n=54. cP<0.01 vs N2.
daf-2(e1370) decreases G93A SOD1-induced paralysis
We next wanted to determine whether increased autophagy in motor neurons of the daf-2(e1370) mutant could protect against G93A SOD1-induced motor neuron dysfunction. We crossed the daf-2(e1370) mutation into the non-GFP fused G93A SOD1 transgenic worms to observe whether autophagy is also induced when G93A SOD1 is expressed in motor neurons of the daf-2(e1370) mutant. Indeed, the number of GFP::LGG-1 puncta in motor neurons of the daf-2(e1370);G93A SOD1 (without the GFP tag) worms was increased, indicating that autophagy is induced in the motor neurons (Figure 2B, 2C). We observed that the daf-2(e1370) mutation significantly decreased G93A SOD1-GFP-induced paralysis (Figure 3C). Additionally, there were more axons that reached the dorsal nerve cord in the daf-2(e1370);G93A worms (Figure 3A, 3D). We therefore asked whether the observed protective effect of the daf-2(e1370) mutation was associated with a reduction of SOD1 aggregations in the transgenic worms. Indeed, we found that in the daf-2(e1370) mutants, the percentage of motor neurons that contained SOD1 aggregations decreased and the SOD1 aggregations also became smaller (Figure 3B, 3E). These results suggest that the daf-2(e1370) mutation protects against the toxic mutant SOD1-induced motor neuron dysfunction by decreasing aggregated/insoluble G93A SOD1.
daf-2(e1370) protects against G93A SOD1-induced motor defect. (A) Confocal image of a daf-2(e1370); G93A worms. Scale bar: 100 μm. (B) G93A SOD1-GFP aggregations in daf-2(e1370); G93A worms. Scale bar: 10 μm. (C) daf-2(e1370) significantly decreases the percentage of paralysis in G93A worms. (D) Quantification the number of axons that reach dorsal nerve cord in daf-2(e1370); G93A and G93A worms. (E) Quantification the percentage of the motor neurons with G93A SOD1 aggregations in daf-2(e1370);G93A and G93A worms. Mean±SEM. n=21. cP<0.01 vs G93A.
Discussion
In this study, we established a C. elegans model of FALS by expressing G93A SOD1 in GABAergic motor neurons. The findings of this study support the hypothesis that protein aggregation is important in mutant SOD1-induced neuron degeneration, during which mutant SOD1 is toxic to C. elegans motor neurons. The damage within the motor neurons alone may be sufficient to produce motor defects. Our findings of the toxicity of mutant SOD1 in C. elegans provides an approach to define the toxic properties of SOD1, specifically in motor neurons, that can lead to motor defects.
Here, we identified the daf-2(e1370) mutant, which is an excellent in vivo genetic model of increased autophagy in C. elegans. Notably, the mRNA level of unc-51 increased much more than the other autophagic genes in daf-2(e1370) mutant. It is known that unc-51, which is the C. elegans ortholog of yeast ATG1, plays an important role in the induction of autophagy. Accordingly, the dramatically increased mRNA levels of unc-51 strongly suggest that autophagy is induced. Previous work has shown an increase in autophagy in hypodermal seam cells that is detectable by visualizing the formation of the autophagosome with GFP::LGG-130. Here, we showed that autophagy is also increased in motor neurons.
Using our C. elegans models of FALS, we found that the daf-2(e1370) mutant is protected from SOD1-induced motor neuron dysfunction. The protective effect of the daf-2(e1370) mutation may be due to reduced aggregation of mutant SOD1 and enhanced recovery of motor neuron function under conditions of mutant SOD1 toxicity. It has been reported that daf-2(e1370) contributes to the suppression of Aβ toxicity expressed in the muscles of C. elegans38. Although we showed here that the daf-2(e1370) mutation could suppress the mutant SOD1-induced toxicity and the autophagy that is increased in the daf-2(e1370) mutant, more direct evidence of the effect of autophagy on toxic SOD1 is also needed. For example, autophagic genes in daf-2(e1370);G93A SOD1 worms could be knocked down to observe whether the protective effect can be suppressed. A forward genetic screen using EMS and a reverse genetics screen using whole-genome RNAi should also be performed in subsequent studies to identify more genes or pathways that participate in mutant SOD1-induced motor dysfunction in C. elegans.
Author contribution
Jia LI and Wei-dong LE designed research; Jia LI performed research; Jia LI and Kai-xing HUANG contributed new reagents or analytic tools; Jia LI and Wei-dong LE analyzed data; Jia LI wrote the paper.
References
Levine B, Kroemer G . Autophagy in the pathogenesis of disease. Cell 2008; 132: 27–42.
Klionsky DJ . Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol 2007; 8: 931–7.
Kroemer G, Levine B . Autophagic cell death: the story of a misnomer. Nat Rev Mol Cell Biol 2008; 9: 1004–10.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451: 1069–75.
Zhang Y, Yan L, Zhou Z, Yang P, Tian E, Zhang K, et al. SEPA-1 mediates the specific recognition and degradation of P granule components by autophagy in C. elegans. Cell 2009; 136: 308–21.
Majumder S, Richardson A, Strong R, Oddo S . Inducing autophagy by rapamycin before, but not after, the formation of plaques and tangles ameliorates cognitive deficits. PLoS One 2011; 6: e25416.
Sarkar S, Perlstein EO, Imarisio S, Pineau S, Cordenier A, Maglathlin RL, et al. Small molecules enhance autophagy and reduce toxicity in Huntington's disease models. Nat Chem Biol 2007; 3: 331–8.
Sarkar S, Ravikumar B, Floto RA, Rubinsztein DC . Rapamycin and mTOR-independent autophagy inducers ameliorate toxicity of polyglutamine-expanded huntingtin and related proteinopathies. Cell Death Differ 2009; 16: 46–56.
Pan T, Kondo S, Zhu W, Xie W, Jankovic J, Le W . Neuroprotection of rapamycin in lactacystin-induced neurodegeneration via autophagy enhancement. Neurobiol Dis 2008; 32: 16–25.
Fornai F, Longone P, Cafaro L, Kastsiuchenka O, Ferrucci M, Manca ML, et al. Lithium delays progression of amyotrophic lateral sclerosis. Proc Natl Acad Sci U S A 2008; 105: 2052–7.
Rowland LP, Shneider NA . Amyotrophic lateral sclerosis. N Engl J Med 2001; 344: 1688–700.
Pasinelli P, Brown RH . Molecular biology of amyotrophic lateral sclerosis: insights from genetics. Nat Rev Neurosci 2006; 7: 710–23.
Rosen DR, Siddique T, Patterson D, Figlewicz DA, Sapp P, Hentati A, et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 1993; 362: 59–62.
Deng HX, Hentati A, Tainer JA, Iqbal Z, Cayabyab A, Hung WY, et al. Amyotrophic lateral sclerosis and structural defects in Cu,Zn superoxide dismutase. Science 1993; 261: 1047–51.
Wroe R, Wai-Ling Butler A, Andersen PM, Powell JF, Al-Chalabi A . ALSOD: the amyotrophic lateral sclerosis online database. Amyotroph Lateral Scler 2008; 9: 249–50.
Valentine JS, Doucette PA, Zittin Potter S . Copper-zinc superoxide dismutase and amyotrophic lateral sclerosis. Annu Rev Biochem 2005; 74: 563–93.
Gurney ME, Pu H, Chiu AY, Dal Canto MC, Polchow CY, Alexander DD, et al. Motor neuron degeneration in mice that express a human Cu, Zn superoxide dismutase mutation. Science 1994; 264: 1772–5.
Wong PC, Pardo CA, Borchelt DR, Lee MK, Copeland NG, Jenkins NA, et al. An adverse property of a familial ALS-linked SOD1 mutation causes motor neuron disease characterized by vacuolar degeneration of mitochondria. Neuron 1995; 14: 1105–16.
Kabuta T, Suzuki Y, Wada K . Degradation of amyotrophic lateral sclerosis-linked mutant Cu,Zn-superoxide dismutase proteins by macroautophagy and the proteasome. J Biol Chem 2006; 281: 30524–33.
Morimoto N, Nagai M, Ohta Y, Miyazaki K, Kurata T, Morimoto M, et al. Increased autophagy in transgenic mice with a G93A mutant SOD1 gene. Brain Res 2007; 1167: 112–7.
Li L, Zhang X, Le W . Altered macroautophagy in the spinal cord of SOD1 mutant mice. Autophagy 2008; 4: 290–3.
Hetz C, Thielen P, Matus S, Nassif M, Court F, Kiffin R, et al. XBP-1 deficiency in the nervous system protects against amyotrophic lateral sclerosis by increasing autophagy. Genes Dev 2009; 23: 2294–306.
Zhang X, Li L, Chen S, Yang D, Wang Y, Wang Z, et al. Rapamycin treatment augments motor neuron degeneration in SOD1(G93A) mouse model of amyotrophic lateral sclerosis. Autophagy 2011; 7: 412–25.
Kuwahara T, Koyama A, Koyama S, Yoshina S, Ren CH, Kato T, et al. A systematic RNAi screen reveals involvement of endocytic pathway in neuronal dysfunction in alpha-synuclein transgenic C. elegans. Hum Mol Genet 2008; 17: 2997–3009.
Cohen E, Bieschke J, Perciavalle RM, Kelly JW, Dillin A . Opposing activities protect against age-onset proteotoxicity. Science 2006; 313: 1604–10.
Silva MC, Fox S, Beam M, Thakkar H, Amaral MD, Morimoto RI . A genetic screening strategy identifies novel regulators of the proteostasis network. PLoS Genet 2011; 7: e1002438.
Wang J, Farr GW, Hall DH, Li F, Furtak K, Dreier L, et al. An ALS-linked mutant SOD1 produces a locomotor defect associated with aggregation and synaptic dysfunction when expressed in neurons of Caenorhabditis elegans. PLoS Genet 2009; 5: e1000350.
Lim MA, Selak MA, Xiang Z, Krainc D, Neve RL, Kraemer BC, et al. Reduced activity of AMP-activated protein kinase protects against genetic models of motor neuron disease. J Neurosci 2012; 32: 1123–41.
Hansen M, Chandra A, Mitic LL, Onken B, Driscoll M, Kenyon C . A role for autophagy in the extension of lifespan by dietary restriction in C. elegans. PLoS Genet 2008; 4: e24.
Melendez A, Talloczy Z, Seaman M, Eskelinen EL, Hall DH, Levine B . Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 2003; 301: 1387–91.
Mello CC, Kramer JM, Stinchcomb D, Ambros V . Efficient gene transfer in C. elegans: extrachromosomal maintenance and integration of transforming sequences. EMBO J 1991; 10: 3959–70.
Brenner S . The genetics of Caenorhabditis elegans. Genetics 1974; 77: 71–94.
Wang J, Farr GW, Zeiss CJ, Rodriguez-Gil DJ, Wilson JH, Furtak K, et al. Progressive aggregation despite chaperone associations of a mutant SOD1-YFP in transgenic mice that develop ALS. Proc Natl Acad Sci U S A 2009; 106: 1392–7.
Witan H, Kern A, Koziollek-Drechsler I, Wade R, Behl C, Clement AM . Heterodimer formation of wild-type and amyotrophic lateral sclerosis-causing mutant Cu/Zn-superoxide dismutase induces toxicity independent of protein aggregation. Hum Mol Genet 2008; 17: 1373–85.
Jonsson PA, Bergemalm D, Andersen PM, Gredal O, Brannstrom T, Marklund SL . Inclusions of amyotrophic lateral sclerosis-linked superoxide dismutase in ventral horns, liver, and kidney. Ann Neurol 2008; 63: 671–5.
Ogura K, Goshima Y . The autophagy-related kinase UNC-51 and its binding partner UNC-14 regulate the subcellular localization of the Netrin receptor UNC-5 in Caenorhabditis elegans. Development 2006; 133: 3441–50.
Melendez A, Levine B . Autophagy in C. elegans. WormBook 2009: Aug 24: 1–26.
Florez-McClure ML, Hohsfield LA, Fonte G, Bealor MT, Link CD . Decreased insulin-receptor signaling promotes the autophagic degradation of beta-amyloid peptide in C. elegans. Autophagy 2007; 3: 569–80.
Acknowledgements
This work was supported by grants from the National Natural Sciences Foundation of China (No 81171201 and No 39070925) and the National Basic Research Program of China (No 2011CB510003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Li, J., Huang, Kx. & Le, Wd. Establishing a novel C. elegans model to investigate the role of autophagy in amyotrophic lateral sclerosis. Acta Pharmacol Sin 34, 644–650 (2013). https://doi.org/10.1038/aps.2012.190
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2012.190
Keywords
This article is cited by
-
From animal models to human disease: a genetic approach for personalized medicine in ALS
Acta Neuropathologica Communications (2016)
-
Mechanisms of aging-related proteinopathies in Caenorhabditis elegans
Experimental & Molecular Medicine (2016)
-
Using C. elegans to discover therapeutic compounds for ageing-associated neurodegenerative diseases
Chemistry Central Journal (2015)
-
Impaired mitochondrial homeostasis and neurodegeneration: towards new therapeutic targets?
Journal of Bioenergetics and Biomembranes (2015)